BootstrapODPSample Variability#
import chainladder as cl
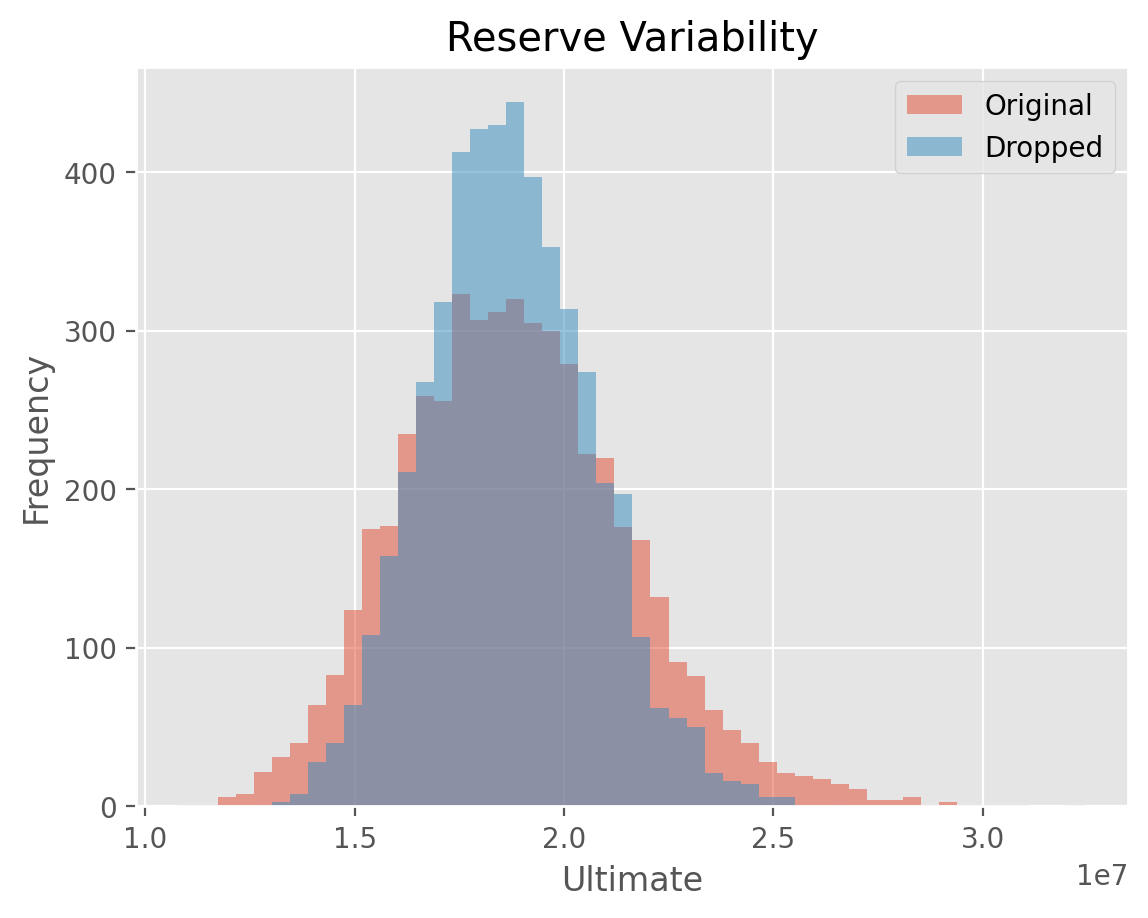
This example demonstrates how you can drop the outlier link ratios from the
BootstrapODPSample. This has a direct consquence of reducing reserve variability estimates.
# Load triangle
triangle = cl.load_sample('genins')
# Use bootstrap sampler to get resampled triangles
s1 = cl.BootstrapODPSample(
n_sims=5000, random_state=42).fit(triangle).resampled_triangles_
## Alternatively use fit_transform() to access resampled triangles dropping
# outlier link-ratios from resampler
s2 = cl.BootstrapODPSample(
drop_high=[True] * 5+ [False] * 4,
drop_low=[True] * 5 + [False] * 4,
n_sims=5000, random_state=42).fit_transform(triangle)
# Summarize results of first model
results = cl.Chainladder().fit(s1).ibnr_.sum('origin').rename('columns', ['Original'])
# Add another column to triangle with second set of results.
results['Dropped'] = cl.Chainladder().fit(s2).ibnr_.sum('origin')
Show code cell source
import matplotlib.pyplot as plt
plt.style.use('ggplot')
%config InlineBackend.figure_format = 'retina'
# Plot both IBNR distributions
ax = results.to_frame().plot(
kind='hist', bins=50, alpha=0.5,
grid=True, xlabel='Ultimate',
title='Reserve Variability')